This post walks through how to create a Siamese network using FastAI. It is based on a tutorial from FastAI. Some of the classes and functions are directly copied from there, but I’ve added things as well.
Table of Contents
First we import everything we need.
import ast
import dill
from fastai.vision.all import *
Then make sure GPU support is available.
torch.cuda.is_available()
True
Data Preparation
We start by downloading the images. We’ll use the cats and dogs dataset because it’s built into FastAI.
path = untar_data(URLs.PETS)
files = get_image_files(path/"images")
Now we’ll need a class to deal with the Siamese images. It’ll make everything easier.
class ImageTuple(fastuple):
@classmethod
def create(cls, file_paths):
"""
creates a tuple of two file paths
"""
return cls(tuple(PILImage.create(f) for f in file_paths))
def show(self, ctx=None, **kwargs):
t1, t2 = self
if not isinstance(t1, Tensor) or not isinstance(t2, Tensor) or t1.shape != t2.shape:
return ctx
line = t1.new_zeros(t1.shape[0], t1.shape[1], 10)
return show_image(torch.cat([t1,line,t2], dim=2), ctx=ctx, **kwargs)
Let’s look at an image.
PILImage.create(files[0])

Now let’s look at a tuple.
img = ImageTuple.create((files[0], files[1]))
tst = ToTensor()(img)
type(tst[0]),type(tst[1])
(fastai.torch_core.TensorImage, fastai.torch_core.TensorImage)
FastAI has a class ToTensor that converts items into tensors. We can use it to visualize the image.
img1 = Resize(224)(img)
tst = ToTensor()(img1)
tst.show();

Now we create an image tuple block. It’s really simple. It just returns a TransformBlock where we pass the function to create a Siamese image.
def ImageTupleBlock():
return TransformBlock(type_tfms=ImageTuple.create, batch_tfms=IntToFloatTensor)
We have to split the data. Let’s do a random split.
splits = RandomSplitter()(files)
splits_files = [files[splits[i]] for i in range(2)]
splits_sets = mapped(set, splits_files)
def get_split(f):
for i,s in enumerate(splits_sets):
if f in s:
return i
raise ValueError(f'File {f} is not presented in any split.')
Now we need a function to return the label given an item. Fortunately, the label is in the filename, so we can use regular expressions to extract it.
def label_func(fname: str):
"""
Extract the label from the file name.
"""
return re.match(r'^(.*)_\d+.jpg$', fname.name).groups()[0]
label_func(files[0])
'havanese'
labels = list(set(files.map(label_func)))
splbl2files = [{l: [f for f in s if label_func(f) == l] for l in labels} for s in splits_sets]
def splitter(items):
"""
This is the function that we actually pass to the DataBlock. All the others are helpers of some form.
"""
def get_split_files(i):
return [j for j,(f1,f2,same) in enumerate(items) if get_split(f1)==i]
return get_split_files(0),get_split_files(1)
def draw_other(f):
"""
Find the pair for the other Siamese image.
"""
same = random.random() < 0.5
cls = label_func(f)
split = get_split(f)
if not same:
cls = random.choice(L(l for l in labels if l != cls))
return random.choice(splbl2files[split][cls]),same
def get_tuples(files):
"""
This function turns a list of files into a list of inputs and label combos.
So each element in this list contains paths of two images and the associated label
"""
return [[f, *draw_other(f)] for f in files]
def get_x(t):
return t[:2]
def get_y(t):
return t[2]
Now we can build the DataBlock.
siamese = DataBlock(
blocks=(ImageTupleBlock, CategoryBlock),
get_items=get_tuples,
get_x=get_x, get_y=get_y,
item_tfms=Resize(224),
batch_tfms=[Normalize.from_stats(*imagenet_stats)]
)
# siamese.summary(files) # long output
Create Dataloaders
Everything looks good. Now we can create our Dataloaders.
dls = siamese.dataloaders(files)
Let’s make sure it looks good.
b = dls.one_batch()
explode_types(b)
{tuple: [{__main__.ImageTuple: [fastai.torch_core.TensorImage,
fastai.torch_core.TensorImage]},
fastai.torch_core.TensorCategory]}
Now we can make a function to show a batch.
@typedispatch
def show_batch(x:ImageTuple, y, samples, ctxs=None, max_n=6, nrows=None, ncols=2, figsize=None, **kwargs):
if figsize is None:
figsize = (ncols*6, max_n//ncols * 3)
if ctxs is None:
ctxs = get_grid(min(len(samples), max_n), nrows=nrows, ncols=ncols, figsize=figsize)
ctxs = show_batch[object](x, y, samples, ctxs=ctxs, max_n=max_n, **kwargs)
return ctxs
dls.show_batch()
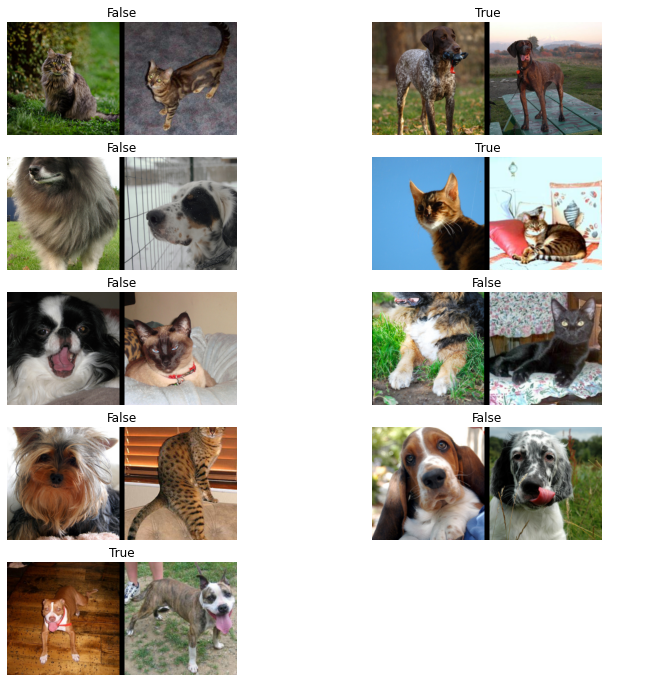

One of my favorite things to do with data in FastAI is turn it into a pandas DataFrame. I find it so much easier to clean or modify the data in the format. In this case, we’ll balance the dataset using a DataFrame.
Convert to DataFrame
def df_label_func(fname):
"""
Extract the label from the data
"""
return fname[2]
def create_dataframe(data, label_func, is_valid=False) -> pd.DataFrame:
"""
Create pandas DataFrame from DataLoaders
"""
items = [x[:2] for x in data.valid.items] if is_valid else [x[:2] for x in data.train.items]
labels = [x[2] for x in data.valid.items] if is_valid else [x[2] for x in data.train.items]
is_valid = [is_valid] * len(items)
return pd.DataFrame({'items': items, 'label': labels, 'is_valid':is_valid})
train_df = create_dataframe(dls, df_label_func, is_valid=False)
valid_df = create_dataframe(dls, df_label_func, is_valid=True)
Let’s see what we have.
train_df.head()
| items | label | is_valid | |
|---|---|---|---|
| 0 | [/home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_159.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_49.jpg] | True | False |
| 1 | [/home/julius/.fastai/data/oxford-iiit-pet/images/leonberger_114.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Russian_Blue_49.jpg] | False | False |
| 2 | [/home/julius/.fastai/data/oxford-iiit-pet/images/beagle_180.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/newfoundland_165.jpg] | False | False |
| 3 | [/home/julius/.fastai/data/oxford-iiit-pet/images/shiba_inu_61.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Abyssinian_103.jpg] | False | False |
| 4 | [/home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_1.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_17.jpg] | True | False |
Now we have it as a dataframe, it’s easier to clean or manipulate the data.
Now let’s oversample the data
train_df['label'].value_counts()
False 2975
True 2937
Name: label, dtype: int64
def oversample(df: pd.DataFrame) -> pd.DataFrame:
"""
We will use pd.DataFrame.sample to oversample from the rows with labels that need to be oversampled.
"""
num_max_labels = df['label'].value_counts().max()
dfs = [df]
for class_index, group in df.groupby('label'):
num_new_samples = num_max_labels - len(group)
dfs.append(group.sample(num_new_samples, replace=True, random_state=42))
return pd.concat(dfs)
oversampled_train_df = oversample(train_df)
oversampled_train_df['label'].value_counts()
True 2975
False 2975
Name: label, dtype: int64
Great. Now we’ll combine it with the validation data again. Note that we didn’t mess with the validation dataset’s label balance.
full_df = pd.concat([oversampled_train_df, valid_df])
Now we make another DataBlock that can read from a csv. Fortunately, FastAI’s support for this is great.
new_dblock = DataBlock(
blocks = (ImageTupleBlock, CategoryBlock),
get_x=ColReader(0),
get_y=ColReader('label'),
splitter = ColSplitter(),
item_tfms = Resize(224),
batch_tfms=[Normalize.from_stats(*imagenet_stats)]
)
new_dls = new_dblock.dataloaders(full_df)
Saving as CSV
If you want to save it as a csv and reload it, you might run into an issue.
full_df.head()
| items | label | is_valid | |
|---|---|---|---|
| 0 | [/home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_159.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_49.jpg] | True | False |
| 1 | [/home/julius/.fastai/data/oxford-iiit-pet/images/leonberger_114.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Russian_Blue_49.jpg] | False | False |
| 2 | [/home/julius/.fastai/data/oxford-iiit-pet/images/beagle_180.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/newfoundland_165.jpg] | False | False |
| 3 | [/home/julius/.fastai/data/oxford-iiit-pet/images/shiba_inu_61.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Abyssinian_103.jpg] | False | False |
| 4 | [/home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_1.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_17.jpg] | True | False |
First, we can’t save the items as Pathlib objects, so we convert them to strings first.
def to_str(x):
return [str(x[0]), str(x[1])]
full_df['items'] = full_df['items'].apply(to_str)
full_df.head()
| items | label | is_valid | |
|---|---|---|---|
| 0 | [/home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_159.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/japanese_chin_49.jpg] | True | False |
| 1 | [/home/julius/.fastai/data/oxford-iiit-pet/images/leonberger_114.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Russian_Blue_49.jpg] | False | False |
| 2 | [/home/julius/.fastai/data/oxford-iiit-pet/images/beagle_180.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/newfoundland_165.jpg] | False | False |
| 3 | [/home/julius/.fastai/data/oxford-iiit-pet/images/shiba_inu_61.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/Abyssinian_103.jpg] | False | False |
| 4 | [/home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_1.jpg, /home/julius/.fastai/data/oxford-iiit-pet/images/pomeranian_17.jpg] | True | False |
Then we can try to save it as a csv.
full_df.to_csv('siamese_data.csv')
full_df2 = pd.read_csv('siamese_data.csv', index_col=0)
full_df2.head()
| items | label | is_valid | |
|---|---|---|---|
| 0 | ['/home/julius/.fastai/data/oxford-iiit-pet/images/beagle_130.jpg', '/home/julius/.fastai/data/oxford-iiit-pet/images/beagle_140.jpg'] | True | False |
| 1 | ['/home/julius/.fastai/data/oxford-iiit-pet/images/havanese_12.jpg', '/home/julius/.fastai/data/oxford-iiit-pet/images/Abyssinian_197.jpg'] | False | False |
| 2 | ['/home/julius/.fastai/data/oxford-iiit-pet/images/leonberger_126.jpg', '/home/julius/.fastai/data/oxford-iiit-pet/images/pug_130.jpg'] | False | False |
| 3 | ['/home/julius/.fastai/data/oxford-iiit-pet/images/saint_bernard_90.jpg', '/home/julius/.fastai/data/oxford-iiit-pet/images/saint_bernard_105.jpg'] | True | False |
| 4 | ['/home/julius/.fastai/data/oxford-iiit-pet/images/american_pit_bull_terrier_123.jpg', '/home/julius/.fastai/data/oxford-iiit-pet/images/keeshond_4.jpg'] | False | False |
Then, if we try to load it, we’ll get an error.
try:
new_dls = new_dblock.dataloaders(full_df2)
except FileNotFoundError as ex:
print(f"This won't work or you'll get this error: {ex}")
This won't work or you'll get this error: [Errno 2] No such file or directory: '['
What you need to do is load the csv into a DataFrame then do a literal_eval. This is because the list has turned into a string representation of a list.
full_df2['items'] = full_df2['items'].apply(ast.literal_eval)
Now it’s good to go.
loaded_dls = new_dblock.dataloaders(full_df2)
next(iter(loaded_dls[0]))
((TensorImage([[[[-1.9638, -1.9980, -2.0152, ..., -1.9124, -1.9124, -1.9124],
[-1.9467, -1.9638, -1.9809, ..., -1.9124, -1.9124, -1.9124],
[-1.9124, -1.9638, -1.9809, ..., -1.9124, -1.9295, -1.9295],
...,
[-2.0152, -2.0152, -1.9809, ..., -2.0152, -1.9809, -1.9809],
[-1.9809, -1.9980, -1.9980, ..., -2.0152, -2.0323, -1.9980],
[-1.9467, -1.9809, -2.0152, ..., -1.9980, -2.0665, -2.0152]],
[[-1.2829, -1.3179, -1.3354, ..., -1.2304, -1.2304, -1.2304],
[-1.2654, -1.2829, -1.3004, ..., -1.2304, -1.2304, -1.2304],
[-1.2304, -1.2829, -1.3004, ..., -1.2304, -1.2479, -1.2479],
...,
[-1.2304, -1.2304, -1.1954, ..., -1.1779, -1.1429, -1.1429],
[-1.1954, -1.2129, -1.2129, ..., -1.1779, -1.1954, -1.1604],
[-1.1604, -1.1954, -1.2304, ..., -1.1604, -1.2304, -1.1779]],
[[-0.8981, -0.9330, -0.9504, ..., -0.8458, -0.8458, -0.8458],
[-0.8807, -0.8981, -0.9156, ..., -0.8458, -0.8458, -0.8458],
[-0.8458, -0.8981, -0.9156, ..., -0.8458, -0.8633, -0.8633],
...,
[-0.8633, -0.8633, -0.8284, ..., -0.8284, -0.7761, -0.7761],
[-0.8284, -0.8458, -0.8458, ..., -0.8284, -0.8458, -0.8110],
[-0.7936, -0.8284, -0.8633, ..., -0.8110, -0.8807, -0.8284]]],
[[[ 0.5193, 0.4508, 0.4851, ..., 0.7248, 0.7077, 0.7248],
[ 0.5364, 0.5022, 0.5022, ..., 0.7248, 0.6906, 0.7077],
[ 0.1254, 0.2624, 0.3481, ..., 0.7419, 0.7248, 0.6906],
...,
[ 0.2282, 0.2453, 0.3138, ..., -0.0972, -0.0629, -0.0458],
[ 0.2796, 0.2967, 0.2796, ..., -0.0972, -0.0801, -0.0458],
[ 0.2796, 0.2967, 0.2796, ..., -0.0972, -0.1143, -0.0972]],
[[ 0.0651, 0.0476, 0.0826, ..., 0.8704, 0.8704, 0.8880],
[ 0.1001, 0.0826, 0.0651, ..., 0.8704, 0.8529, 0.8704],
[-0.2850, -0.1275, -0.0224, ..., 0.8880, 0.8704, 0.8529],
...,
[ 0.6429, 0.6779, 0.6604, ..., 0.2752, 0.2927, 0.3102],
[ 0.6429, 0.6604, 0.6779, ..., 0.2577, 0.2752, 0.2927],
[ 0.6779, 0.6604, 0.6779, ..., 0.2402, 0.2402, 0.2402]],
[[ 0.9494, 0.9494, 0.9494, ..., 1.1062, 1.1237, 1.1934],
[ 0.9668, 0.9842, 0.9668, ..., 1.0888, 1.1062, 1.1411],
[ 0.5485, 0.7054, 0.8099, ..., 1.1237, 1.1237, 1.1062],
...,
[ 0.6705, 0.6879, 0.7054, ..., 0.3393, 0.3393, 0.3568],
[ 0.6879, 0.7054, 0.7054, ..., 0.3219, 0.3219, 0.3393],
[ 0.7228, 0.7054, 0.7054, ..., 0.3219, 0.2871, 0.2871]]],
[[[ 0.1254, 0.0912, 0.0741, ..., -1.2788, -0.5938, 1.5468],
[ 0.1597, 0.1254, 0.1254, ..., -1.3130, -0.6109, 1.5297],
[ 0.1768, 0.1597, 0.1597, ..., -1.2788, -0.6452, 1.4954],
...,
[ 1.6838, 1.7180, 1.7352, ..., 0.8961, 1.5297, 1.9407],
[ 2.0777, 2.0948, 2.0948, ..., 1.8893, 1.9064, 2.0263],
[ 2.0434, 2.0605, 2.0777, ..., 2.0263, 2.0434, 2.0948]],
[[-0.8452, -0.8803, -0.8978, ..., -1.3179, -0.6176, 1.5532],
[-0.8102, -0.8452, -0.8452, ..., -1.3529, -0.6352, 1.5357],
[-0.7927, -0.8102, -0.8102, ..., -1.3179, -0.6702, 1.5007],
...,
[ 1.4832, 1.5182, 1.5357, ..., 0.7479, 1.4657, 1.9384],
[ 1.9734, 2.0084, 2.0084, ..., 1.7633, 1.8683, 2.0259],
[ 2.0434, 2.0609, 2.0784, ..., 1.9734, 2.0259, 2.0784]],
[[-1.3339, -1.3687, -1.3861, ..., -1.3164, -0.6018, 1.6117],
[-1.2990, -1.3339, -1.3339, ..., -1.3513, -0.6193, 1.5942],
[-1.2816, -1.2990, -1.2990, ..., -1.3164, -0.6367, 1.5594],
...,
[ 1.1934, 1.2282, 1.2457, ..., 0.7228, 1.4897, 1.9777],
[ 2.1171, 2.1346, 2.1520, ..., 1.9080, 2.0300, 2.2043],
[ 2.4134, 2.4309, 2.4483, ..., 2.2391, 2.3088, 2.3786]]],
...,
[[[-2.1008, -2.0837, -2.0837, ..., 0.5364, 1.4098, 1.7865],
[-2.1008, -2.0837, -2.0837, ..., 0.7591, 1.5125, 1.8379],
[-2.1008, -2.0837, -2.0837, ..., 0.9474, 1.6324, 1.8893],
...,
[-2.0494, -2.0152, -2.0323, ..., -1.1075, -0.8164, 0.0398],
[-2.0494, -2.0494, -2.0665, ..., -0.3027, 0.7419, 1.6495],
[-2.0665, -2.0665, -2.0494, ..., 0.5536, 1.9749, 2.2147]],
[[-2.0182, -2.0007, -2.0007, ..., 0.6604, 1.5357, 1.9209],
[-2.0182, -2.0007, -2.0007, ..., 0.8704, 1.6583, 1.9734],
[-2.0182, -2.0007, -2.0007, ..., 1.0805, 1.7808, 2.0259],
...,
[-1.9482, -1.9307, -1.9482, ..., -1.1429, -1.1954, -0.6352],
[-1.9307, -1.9307, -1.9482, ..., -1.1429, -0.4776, 0.4503],
[-1.9482, -1.9482, -1.9307, ..., 0.1001, 1.8158, 2.3761]],
[[-1.7870, -1.7696, -1.7696, ..., 0.1825, 0.9842, 1.4374],
[-1.7870, -1.7696, -1.7696, ..., 0.3568, 1.0714, 1.4897],
[-1.7870, -1.7696, -1.7696, ..., 0.5485, 1.2108, 1.5420],
...,
[-1.7347, -1.6999, -1.7173, ..., -0.9156, -0.8981, -0.2358],
[-1.7173, -1.7173, -1.7347, ..., -0.7064, 0.0431, 1.0017],
[-1.7347, -1.7347, -1.7173, ..., 0.5659, 2.1171, 2.5703]]],
[[[-2.0665, -2.0494, -2.0494, ..., -2.0837, -2.0665, -2.0837],
[-0.5767, -0.5767, -0.5767, ..., -0.5596, -0.5424, -0.5938],
[ 1.5810, 1.5982, 1.3584, ..., 1.6153, 1.5810, 1.5810],
...,
[ 1.4612, 1.5125, 1.5125, ..., 1.4954, 1.4954, 1.5468],
[-0.5767, -0.5767, -0.5938, ..., -0.5596, -0.5596, -0.5424],
[-2.0494, -2.0665, -2.0837, ..., -2.0837, -2.1008, -2.0494]],
[[-1.9832, -1.9657, -1.9832, ..., -1.9657, -1.9657, -2.0007],
[-0.4426, -0.4251, -0.4601, ..., -0.4601, -0.4426, -0.4601],
[ 1.7283, 1.7283, 1.4832, ..., 1.7108, 1.6933, 1.7108],
...,
[ 1.5532, 1.5882, 1.6232, ..., 1.5182, 1.5007, 1.5532],
[-0.4601, -0.4426, -0.4426, ..., -0.4426, -0.4426, -0.4076],
[-2.0007, -1.9832, -2.0007, ..., -2.0007, -2.0007, -1.9657]],
[[-1.7173, -1.7522, -1.7696, ..., -1.7522, -1.7173, -1.7522],
[-0.2532, -0.2358, -0.2184, ..., -0.2532, -0.2358, -0.2184],
[ 1.7337, 1.7685, 1.5420, ..., 1.8208, 1.8034, 1.8034],
...,
[ 1.5768, 1.6465, 1.6814, ..., 1.5942, 1.5942, 1.6465],
[-0.2532, -0.2184, -0.2184, ..., -0.2184, -0.2184, -0.1835],
[-1.7347, -1.7870, -1.7870, ..., -1.7522, -1.7522, -1.6824]]],
[[[ 0.2282, 0.1939, 0.1254, ..., -0.3712, -0.3712, -0.3712],
[ 0.2453, 0.1768, 0.0741, ..., -0.3712, -0.3712, -0.3712],
[ 0.2111, 0.1597, 0.0912, ..., -0.3712, -0.3712, -0.3712],
...,
[-0.7993, -0.8678, -0.9020, ..., -0.3883, -0.4226, -0.3883],
[-0.6623, -0.4911, -0.5938, ..., -0.0629, -0.2684, -0.4911],
[-0.5082, -0.7137, -0.9192, ..., -0.5424, -0.5767, -0.7479]],
[[ 0.1877, 0.1527, 0.1001, ..., -0.3200, -0.3200, -0.3200],
[ 0.2052, 0.1352, 0.0476, ..., -0.3200, -0.3200, -0.3200],
[ 0.1702, 0.1352, 0.0651, ..., -0.3200, -0.3200, -0.3200],
...,
[-0.5301, -0.6001, -0.6352, ..., -0.1625, -0.1975, -0.1625],
[-0.3901, -0.2150, -0.3200, ..., 0.1702, -0.0399, -0.2675],
[-0.2850, -0.4951, -0.7227, ..., -0.3725, -0.4076, -0.5826]],
[[ 0.2522, 0.2173, 0.1651, ..., -0.2881, -0.2881, -0.2881],
[ 0.2696, 0.1999, 0.1128, ..., -0.2881, -0.2881, -0.2881],
[ 0.2348, 0.1999, 0.1302, ..., -0.2881, -0.2881, -0.2881],
...,
[-0.3927, -0.4624, -0.4973, ..., -0.1487, -0.1835, -0.1487],
[-0.2532, -0.0790, -0.1835, ..., 0.1825, -0.0267, -0.2532],
[-0.1312, -0.3404, -0.5670, ..., -0.3055, -0.3404, -0.5147]]]],
device='cuda:0'),
TensorImage([[[[-1.3130, -1.3130, -1.3302, ..., -1.8268, -1.8268, -1.8268],
[-1.2959, -1.2959, -1.3130, ..., -1.8268, -1.8268, -1.8439],
[-1.2959, -1.2959, -1.3130, ..., -1.8268, -1.8268, -1.8610],
...,
[ 1.0502, 0.9817, 0.9303, ..., -0.5767, -0.4397, -0.2684],
[ 1.2043, 1.1872, 1.1529, ..., -0.6281, -0.3541, -0.3198],
[ 1.3927, 1.4098, 1.3584, ..., -0.4911, -0.3883, -0.2171]],
[[-1.2129, -1.2129, -1.2304, ..., -1.7381, -1.7381, -1.7381],
[-1.1954, -1.1954, -1.2129, ..., -1.7381, -1.7381, -1.7556],
[-1.1954, -1.1954, -1.2129, ..., -1.7381, -1.7381, -1.7731],
...,
[ 1.1856, 1.0805, 1.0105, ..., -0.6877, -0.5476, -0.3725],
[ 1.3431, 1.2906, 1.2731, ..., -0.7402, -0.4776, -0.4426],
[ 1.5357, 1.5357, 1.4832, ..., -0.5651, -0.4601, -0.2850]],
[[-0.9853, -0.9853, -1.0027, ..., -1.4733, -1.4733, -1.4733],
[-0.9678, -0.9678, -0.9853, ..., -1.4733, -1.4733, -1.4907],
[-0.9678, -0.9678, -0.9853, ..., -1.4733, -1.4733, -1.5081],
...,
[ 1.3677, 1.2457, 1.1585, ..., -0.5844, -0.4450, -0.2707],
[ 1.5245, 1.4897, 1.4200, ..., -0.6541, -0.3753, -0.3578],
[ 1.7337, 1.7685, 1.6814, ..., -0.4798, -0.3927, -0.2184]]],
[[[ 1.7523, 1.7523, 1.7523, ..., 0.4679, 0.4679, 0.4679],
[ 1.7523, 1.7523, 1.7523, ..., 0.4679, 0.4679, 0.4679],
[ 1.7523, 1.7523, 1.7523, ..., 0.4679, 0.4679, 0.4679],
...,
[ 0.6221, 0.5193, 0.5364, ..., -0.1828, -0.2342, -0.1657],
[ 0.5364, 0.4337, 0.5022, ..., -0.1657, -0.2513, -0.1657],
[ 0.4508, 0.3481, 0.4508, ..., -0.1657, -0.2513, -0.1657]],
[[ 1.9209, 1.9209, 1.9209, ..., 0.4678, 0.4678, 0.4678],
[ 1.9209, 1.9209, 1.9209, ..., 0.4678, 0.4678, 0.4678],
[ 1.9209, 1.9209, 1.9209, ..., 0.4678, 0.4678, 0.4678],
...,
[ 0.5903, 0.4853, 0.4503, ..., -0.3725, -0.3375, -0.2500],
[ 0.5028, 0.3803, 0.4153, ..., -0.3550, -0.3550, -0.2500],
[ 0.4153, 0.2927, 0.3627, ..., -0.3550, -0.3550, -0.2500]],
[[ 2.0997, 2.0997, 2.0997, ..., 0.4962, 0.4962, 0.4962],
[ 2.0997, 2.0997, 2.0997, ..., 0.4962, 0.4962, 0.4962],
[ 2.0997, 2.0997, 2.0997, ..., 0.4962, 0.4962, 0.4962],
...,
[ 0.2348, 0.0953, 0.0779, ..., -0.4973, -0.4275, -0.3230],
[ 0.1476, -0.0092, 0.0256, ..., -0.4798, -0.4450, -0.3230],
[ 0.0605, -0.0964, -0.0092, ..., -0.4798, -0.4450, -0.3230]]],
[[[ 0.5878, 0.7077, 0.8104, ..., -1.7069, -1.6555, -1.7583],
[ 0.5022, 0.5022, 0.4851, ..., -1.6727, -1.7754, -1.8268],
[ 0.3823, 0.2967, 0.3652, ..., -1.8268, -1.7925, -1.8439],
...,
[ 0.0741, 0.1083, 0.1083, ..., -0.5424, -0.4739, -0.4397],
[ 0.1083, 0.1426, 0.0398, ..., -0.6109, -0.4397, -0.5938],
[ 0.0398, 0.0741, 0.0056, ..., -0.5596, -0.6281, -0.4739]],
[[-0.5301, -0.1275, -0.0049, ..., -1.8431, -1.8606, -1.8081],
[-0.2150, -0.0399, -0.0224, ..., -1.8606, -1.7906, -1.8081],
[ 0.0651, 0.0826, 0.0651, ..., -1.7731, -1.7731, -1.7906],
...,
[-0.5826, -0.5476, -0.5126, ..., -1.0553, -0.8627, -1.0378],
[-0.6527, -0.6527, -0.6877, ..., -1.0903, -0.9328, -1.1429],
[-0.4951, -0.4951, -0.4951, ..., -1.3179, -1.1604, -0.9853]],
[[-1.5430, -1.1596, -1.0898, ..., -1.5953, -1.6824, -1.6476],
[-0.9330, -0.6193, -0.6367, ..., -1.5953, -1.7173, -1.7522],
[-0.2184, -0.2184, -0.2358, ..., -1.7522, -1.7522, -1.7347],
...,
[-1.0898, -1.0550, -0.9853, ..., -1.3513, -1.2641, -1.3513],
[-1.1247, -1.1421, -1.1596, ..., -1.3513, -1.2990, -1.3861],
[-1.1944, -1.1944, -1.1770, ..., -1.2990, -1.2816, -1.2816]]],
...,
[[[-1.6213, -1.6898, -1.7069, ..., -2.0323, -1.9980, -1.9467],
[-1.6555, -1.7240, -1.7240, ..., -2.0494, -2.0152, -1.9124],
[-1.6727, -1.7412, -1.7412, ..., -2.0665, -2.0152, -1.8953],
...,
[-1.7412, -1.7754, -1.8439, ..., -2.0665, -2.0837, -2.1008],
[-1.7925, -1.8439, -1.9124, ..., -2.1179, -2.1179, -2.1179],
[-1.7412, -1.7925, -1.8610, ..., -2.1179, -2.1179, -2.1179]],
[[-0.3025, -0.3550, -0.3550, ..., -0.5826, -0.5476, -0.4951],
[-0.3375, -0.3901, -0.3725, ..., -0.6001, -0.5651, -0.4601],
[-0.3550, -0.4076, -0.3901, ..., -0.6176, -0.5651, -0.4426],
...,
[-1.0378, -1.0728, -1.0903, ..., -1.2129, -1.2304, -1.2479],
[-1.0378, -1.0728, -1.0903, ..., -1.1779, -1.1954, -1.2129],
[-0.9153, -0.9328, -0.9503, ..., -1.0903, -1.1078, -1.1078]],
[[-0.4973, -0.5495, -0.5495, ..., -0.8110, -0.7761, -0.7238],
[-0.5321, -0.5844, -0.5670, ..., -0.8284, -0.7936, -0.6890],
[-0.5495, -0.6018, -0.5844, ..., -0.8458, -0.7936, -0.6715],
...,
[-1.0376, -1.0724, -1.1073, ..., -1.3164, -1.3339, -1.3513],
[-1.0376, -1.0724, -1.1073, ..., -1.3339, -1.3513, -1.3513],
[-0.9330, -0.9678, -1.0027, ..., -1.2816, -1.2990, -1.2990]]],
[[[ 0.5878, 0.4508, 0.1254, ..., -0.1999, -0.0801, 0.3652],
[ 0.3481, 0.1254, 0.1768, ..., 0.2967, -0.0801, 0.2624],
[ 0.2453, 0.0398, 0.1939, ..., 0.4851, 0.1254, 0.3309],
...,
[-0.4054, -0.1999, -0.2171, ..., 1.6153, 1.6324, 1.7352],
[-0.2342, -0.0972, -0.1486, ..., 1.7180, 1.5468, 1.7352],
[-0.2171, -0.1314, 0.0398, ..., 1.7865, 1.6495, 1.6667]],
[[ 0.9055, 0.7654, 0.4328, ..., 0.1176, 0.2402, 0.6954],
[ 0.6604, 0.4328, 0.4853, ..., 0.6254, 0.2402, 0.5903],
[ 0.5553, 0.3452, 0.5028, ..., 0.8179, 0.4503, 0.6604],
...,
[-0.0749, 0.1352, 0.1176, ..., 2.1310, 2.2535, 2.3585],
[ 0.1001, 0.2402, 0.1877, ..., 2.2360, 2.1660, 2.3585],
[-0.0574, 0.0826, 0.2927, ..., 2.3235, 2.2535, 2.2710]],
[[ 1.1062, 0.9668, 0.6356, ..., 0.2348, 0.3916, 0.8448],
[ 0.8622, 0.6356, 0.6879, ..., 0.7402, 0.3916, 0.7402],
[ 0.7576, 0.5485, 0.7054, ..., 0.9319, 0.6008, 0.8099],
...,
[ 0.0779, 0.2871, 0.2696, ..., 2.4657, 2.4831, 2.5877],
[ 0.2522, 0.3916, 0.3393, ..., 2.5703, 2.3960, 2.5877],
[ 0.0779, 0.1999, 0.3916, ..., 2.5877, 2.5529, 2.5703]]],
[[[ 2.2489, 2.2489, 2.2489, ..., 1.2385, 1.2385, 1.2214],
[ 2.2489, 2.2489, 2.2489, ..., 1.2557, 1.2385, 1.2214],
[ 2.2489, 2.2489, 2.2489, ..., 1.2557, 1.2214, 1.2043],
...,
[ 1.0331, 0.9817, 0.9303, ..., -1.1760, -1.3644, -1.5357],
[ 0.6906, 0.6734, 0.6392, ..., -1.1247, -1.3130, -1.4843],
[ 0.4508, 0.4166, 0.3994, ..., -1.0390, -1.2445, -1.4329]],
[[ 2.4286, 2.4286, 2.4286, ..., -0.0749, -0.0924, -0.1099],
[ 2.4286, 2.4286, 2.4286, ..., -0.0399, -0.0574, -0.0749],
[ 2.4286, 2.4286, 2.4286, ..., -0.0574, -0.0399, -0.0749],
...,
[ 0.3627, 0.3277, 0.2752, ..., -1.3880, -1.5105, -1.6331],
[-0.0049, -0.0224, -0.0749, ..., -1.3354, -1.4580, -1.5980],
[-0.2850, -0.3200, -0.3550, ..., -1.3004, -1.4230, -1.5630]],
[[ 2.6400, 2.6400, 2.6400, ..., -1.3164, -1.3687, -1.3339],
[ 2.6400, 2.6400, 2.6400, ..., -1.3339, -1.3513, -1.3339],
[ 2.6400, 2.6400, 2.6400, ..., -1.3513, -1.3687, -1.3513],
...,
[ 0.8448, 0.8099, 0.7402, ..., -1.2816, -1.4036, -1.5081],
[ 0.4788, 0.4439, 0.3916, ..., -1.2641, -1.3861, -1.5081],
[ 0.1825, 0.1302, 0.0779, ..., -1.2467, -1.4036, -1.4907]]]],
device='cuda:0')),
TensorCategory([0, 1, 1, 1, 1, 1, 0, 1, 0, 1, 0, 1, 0, 1, 0, 0, 1, 0, 1, 1, 1, 1, 0, 0,
0, 0, 0, 0, 1, 0, 1, 1, 1, 0, 1, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0, 1, 1, 0,
0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 0, 1, 1, 0], device='cuda:0'))
Modeling
Now we build a simple Siamese Model.
class SiameseModel(Module):
def __init__(self, encoder, head):
self.encoder, self.head = encoder, head
def forward(self, x):
"""
This takes x1 and x2 as two separate things. But if we change it to x, maybe we're OK?
"""
x1, x2 = x
ftrs = torch.cat([self.encoder(x1), self.encoder(x2)], dim=1)
return self.head(ftrs)
model_meta[resnet34]
{'cut': -2,
'split': <function fastai.vision.learner._resnet_split(m)>,
'stats': ([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])}
encoder = create_body(resnet34, cut=-2)
head = create_head(512*2, 2, ps=0.5)
model = SiameseModel(encoder, head)
def siamese_splitter(model):
return [params(model.encoder), params(model.head)]
def loss_func(out, targ):
return CrossEntropyLossFlat()(out, targ.long())
learn = Learner(loaded_dls, model, loss_func=loss_func, splitter=siamese_splitter, metrics=accuracy)
learn.freeze()
learn.lr_find()
/home/julius/miniconda3/envs/pt/lib/python3.8/site-packages/torch/nn/functional.py:718: UserWarning: Named tensors and all their associated APIs are an experimental feature and subject to change. Please do not use them for anything important until they are released as stable. (Triggered internally at /opt/conda/conda-bld/pytorch_1623448234945/work/c10/core/TensorImpl.h:1156.)
return torch.max_pool2d(input, kernel_size, stride, padding, dilation, ceil_mode)
SuggestedLRs(valley=0.0012022644514217973)
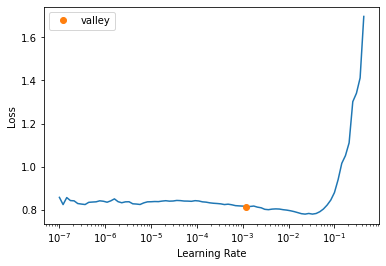
learn.fit_one_cycle(4, 1e-3)
| epoch | train_loss | valid_loss | accuracy | time |
|---|---|---|---|---|
| 0 | 0.599453 | 0.396537 | 0.820704 | 00:23 |
| 1 | 0.362100 | 0.271485 | 0.895805 | 00:23 |
| 2 | 0.243444 | 0.228767 | 0.914750 | 00:23 |
| 3 | 0.176664 | 0.220484 | 0.917456 | 00:23 |
There you go! A fully trained Siamese network.
Save Model
Now let’s save it.
learn.path = Path(os.getenv('MODELS'))
learn.export('siamese_catsvdogs.pkl', pickle_module=dill)